Quantitative Histology
Image processing and analyses are today’s invaluable tools to translate pathology into numbers. Our philosophy is to create high-throughput procedures that are as rater independent as possible and replicable at every step to data.
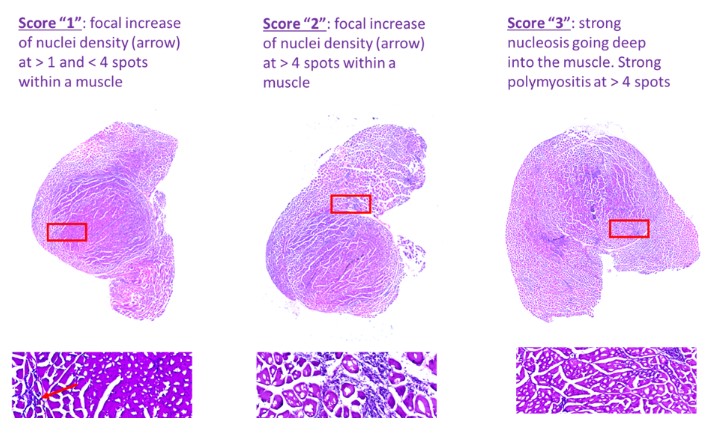
PGI can offer a wide range of different kind of measurements. It starts with simple semi-quantitative rating (e.g. Figure 1) for muscle infiltration scoring, includes a plethora of fully and semi-automated standard measurements (e.g. Figure 2) for determination of labeled surface area, and reaches to complex computer-learned segmentation of classified objects (e.g. Figure 3) for microglia classification.
Our goal is full comprehensibility of measurements. Therefore, in-house macroprogramming serves to automatically store all measurement parameters together with the measurement, as well as the result masks to be able to visualize and reproduce every count.
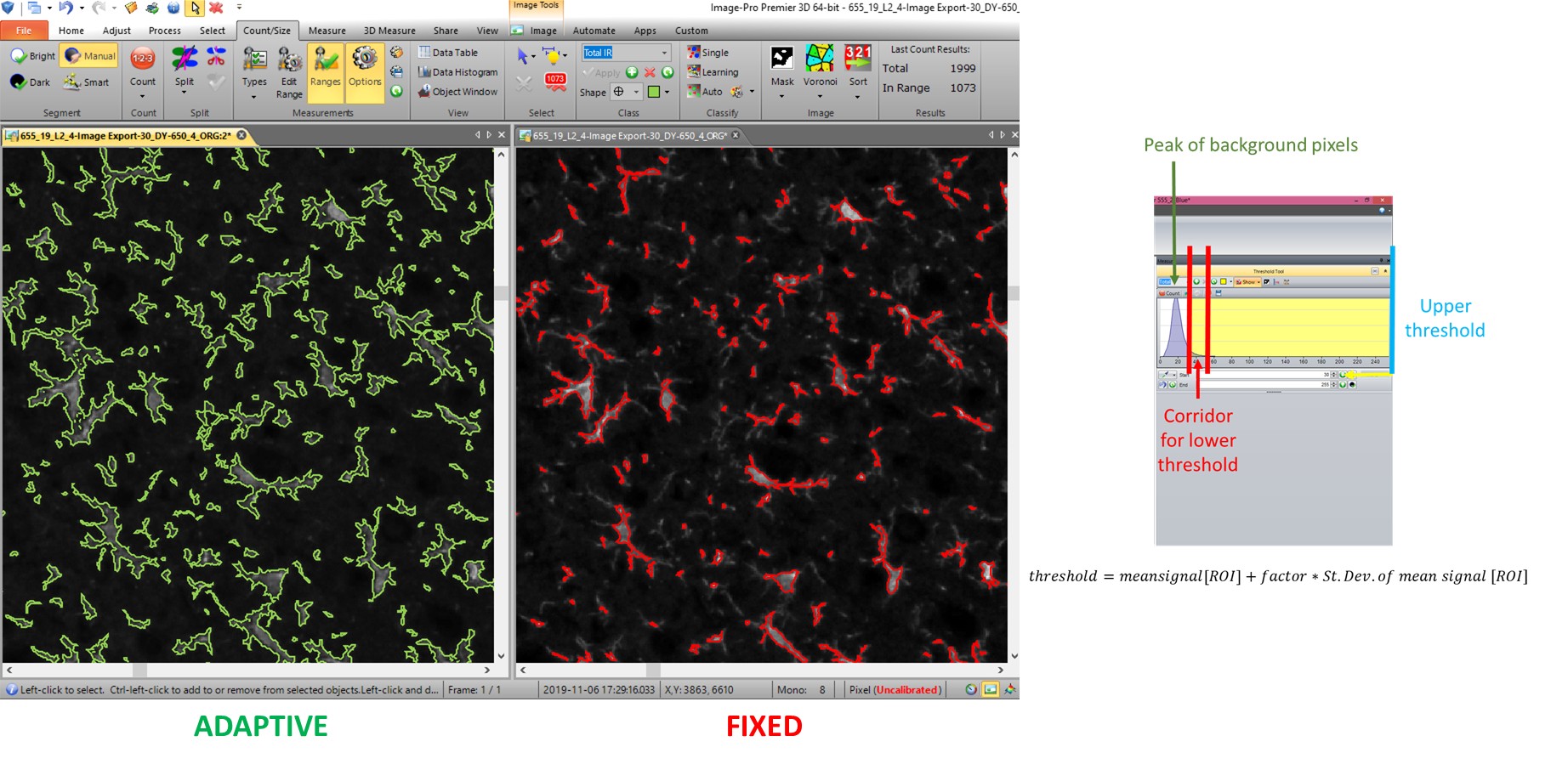
The macros allow the use of complex procedures like filtering, adaptive thresholding and image arithmetics to facilitate uniform object detection. For instance, classical thresholds for object detection are intensity based fixed numbers and thus do not segment objects in accordance with their real physical borders but present signal on the object. They are vulnerable to textural bias and measurements in different groups can lead to measurements of different kind of objects. Limited adaptive thresholds as shown in Figure 4 cope with this infirmity.
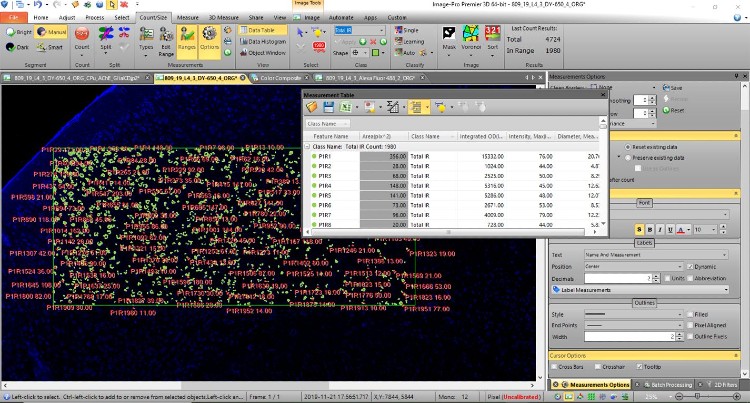
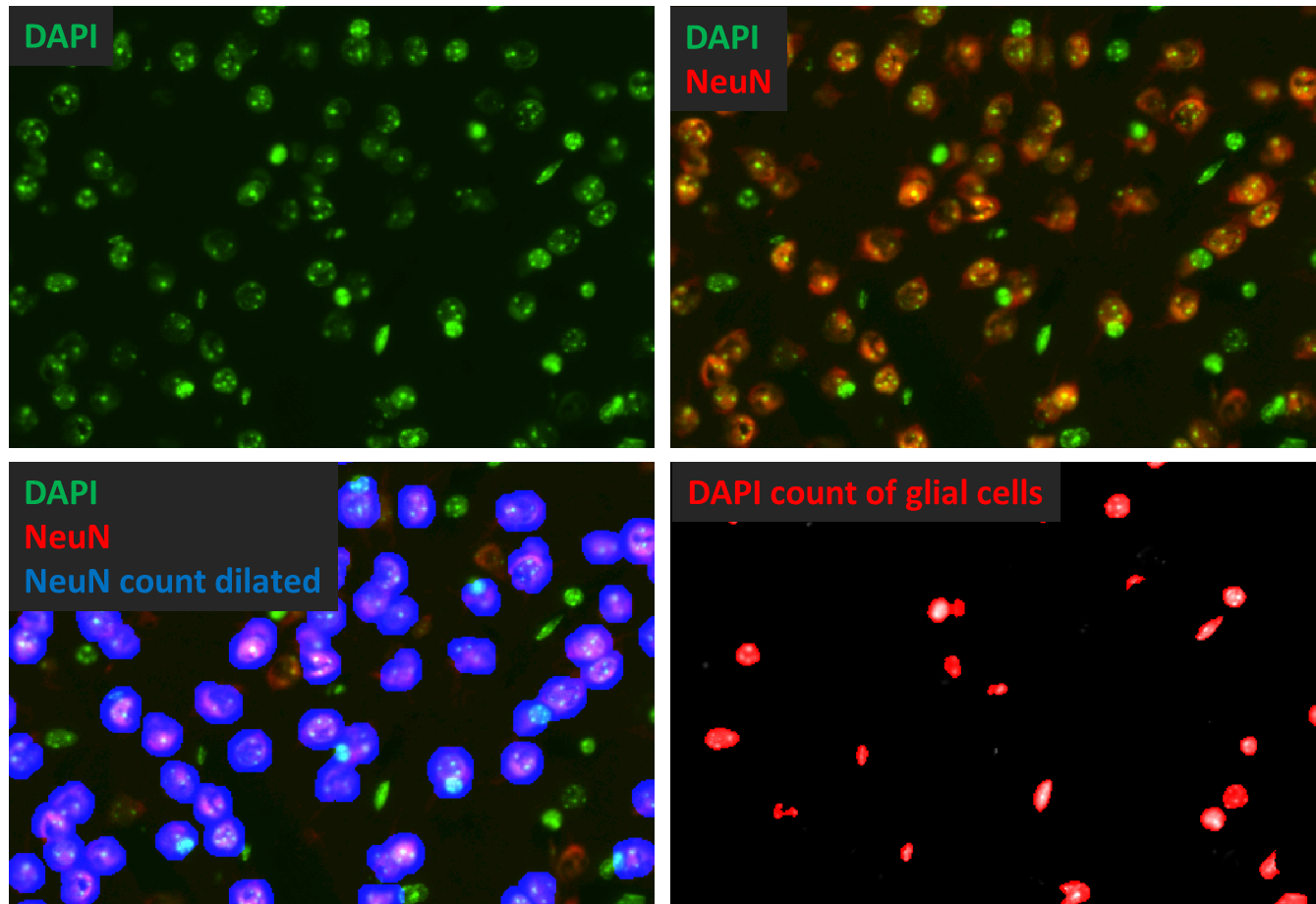
Figure 5 shows an example for image arithmetics and the use of measurement masks to extract specific information. In this case a separation of neuronal and glial DAPI nuclei is done by masking all neuronal nuclei before the count of DAPI.
Such nuclei masks can be stored and used to debias cell counts of all kind of cells . The latter combined with certain object detection parameters (fill holes, pixel connect options) assure that exclusively cells are counted that have a full nucleus within the section (reduction of 3D bias). An example of a count is shown in Figure 6.
Those few examples can only show a small portion of our capabilities. We are also happy to develop, customize and automate your personal evaluations on a variety of image analyses platforms.
Figure 1: Example of semi-quantitative scoring of blood cell infiltration in gastrocnemius muscle of mdx mice. Scoring system for semi-quantitative analysis of muscle inflammation. Note that an additional score was added to increase separation sensitivity of the score suiting to the evident muscle pathology in the samples. Score “0’ is healthy and not shown here.
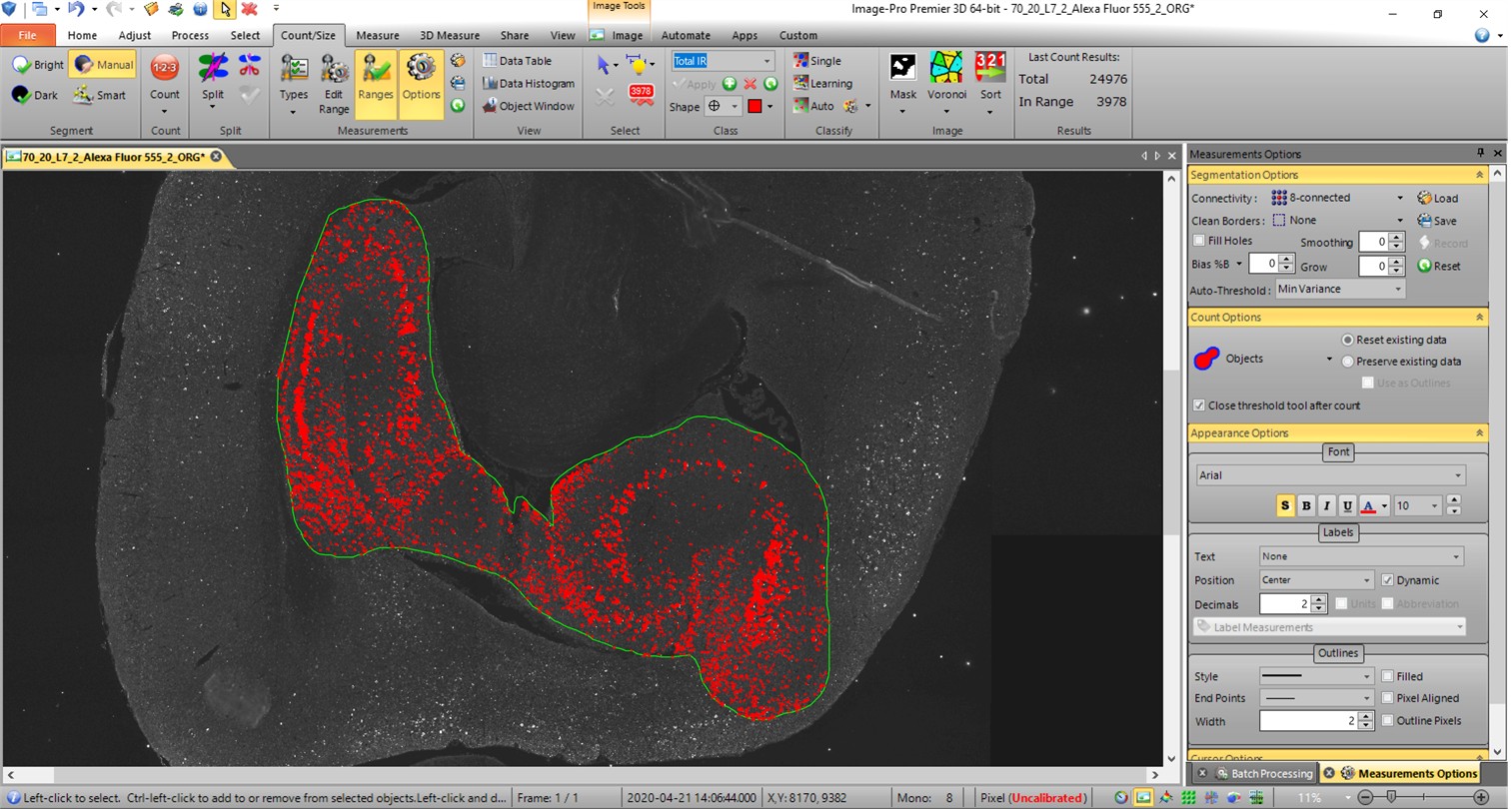
Figure 2: Example of determination of IR area percentage of an immuno-labeling (AT8) in the hippocampus within green manually delineated outline in a model of tauopathy (rTg4510).
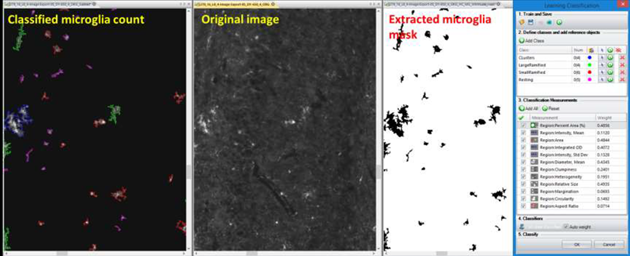
Figure 3: Example of microglial cell classification with computer learned algorithms to separate large and small ramifying microglia from ramified resting state and cell clusters.
Figure 4: Example of limited adaptive thresholding in whole slice images versus classical fixed intensity threshold. The threshold is determined by a formula that accounts for individual histogram variation on the level of single sections. In combination with upper and lower threshold limits the adaptive thresholding guarantees a more uniform detection of structures than classical fixed intensity based thresholds that are vulnerable to over-, or as in this case, underestimation of real object size.
Figure 5: Example of image arithmetics to separate neuronal from glial nuclei: A measurement mask of NeuNpositive neuronal nuclei (neuronal nuclei primary antibody, here shown in blue pseudo tint in Lower Right) is stored, dilated and subtracted from the original DAPI image (green). The image in which all neuronal nuclei are veiled is shown in the lower right corner. The resulting count (red) reflects glial nuclei only.
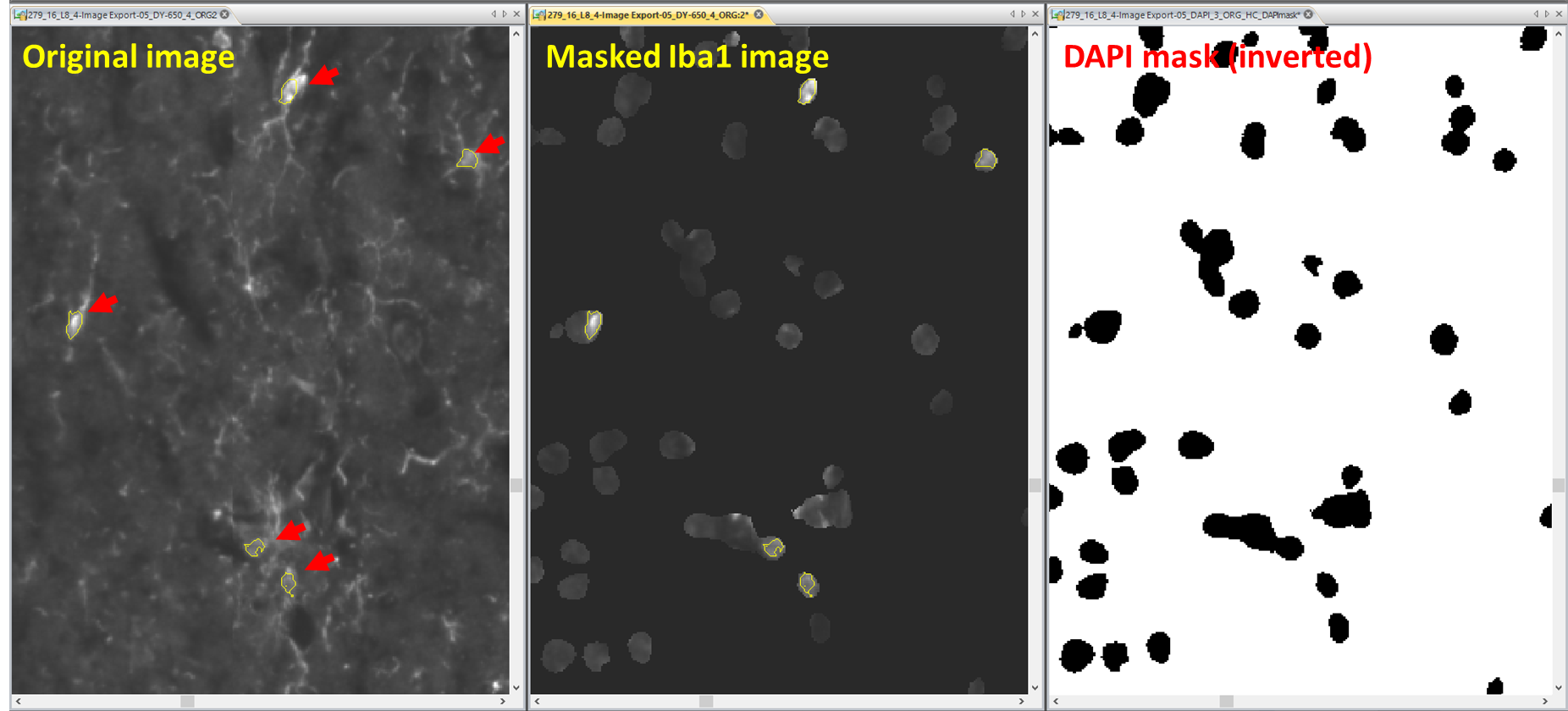
Figure 6: Debiasing cell density for e.g. microbiological cells: To determine the number of cells that have their soma within the section level, an inverted bi-level (black & white) mask of the DAPI count (Right Panel) is subtracted from the lba1 image (Middle Panel). Nuclei overlapping with lba1 signal are counted using pixel connect and fill holes options for detection. Overlaying the count of the original image (Left Panel) shows that only lba1 positive cells are counted if they have a nucleus within the section level (marked by arrows), while all other objects (processes, truncated somata without full nucleus) are not counted. This count related to ROI size estimates the cell destiny very precisely.